Cell Line Derived Xenografts (CDX). Over 60 xenograft models available at Altogen Labs.
Cell Line Derived Xenograft models (CDX models) provide decision-making checkpoints that gate whether a test agent moves forward in the pipeline. CDX models allow clients to further advance preclinical development programs optimally and in a budget conscious means. Mouse CDX xenografts are created by injecting tumor cells from one species into a different species (i.e. human cancer cell lines are inoculated into immune compromised mice). There are many CROs that offer Xenograft services, but knowledge and experience using the various models is a key factor in experimental success. In vivo transfection reagents are used to determine tumor growth delay or tumor growth inhibition in animal models.
| Tumor Type | Cell Line |
| Brain | LN-229, SF268, SF295, SF539, SK-N-AS, SNB-19, SNB-75, U87 MG, U-251 MG |
| Breast | BT-474, HS578T, KPL-4, MCF-7, MDA-MB-231, MDA-MB-453, T-47D |
| Colon | COLO 205, DLD-1, HCT 116, HT-29, LoVo, LS174T, RKO, SW-620, WiDr |
| Lung | A549, Calu-3, H1155, H460, NCI-H1975, NCI-H226 |
| Melanoma | A375, A431, A2058, B16, SK-MEL-2 |
| Hematological | HL60, K562, DOHH2, KARPAS 299, Raji, Ramos |
| Pancreas | BxPC-3, MIA PaCa-2, PANC-1 |
* Table courtesy of Altogen Labs
List of citations and references for Altogen Transfection Services and Products
In Vivo Imaging
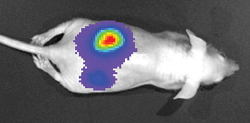
Subcutaneous, orthotopic and metastatic Cell Line Derived Xenograft models that express a bioluminescent signal enables researchers to capture optical images. These models provide information for diseases at different stages, with captured images of the primary, spontaneous and experimental metastases replicating the progression of early stage to late stage disease.
The use of metastatic, cell line derived xenograft animal models allow the researcher to track anti- tumor activity internally at a metastatic region, or perhaps in the cells naturally occurring tissue environment (i.e. liver tumor cells implanted in the mouse liver). In vivo transfection in xenograft tumor models is an established method for testing efficacy of a test compound while considering tumor microenvironment. For this, cells that express exogenous genes such as luciferase, Green Fluorescent Protein (GFP) or Red Fluorescent Protein (RFP) are used to track tumor growth delay during progression of… [Read More]
Liposome Encapsulation
Liposomes provide a protective barrier for the nucleic acid cargo in the internal aqueous core (e.g. DNA, siRNA, miRNA) for both in vitro and in vivo transfection experiments. This includes standard transfection formulations (i.e. cholesterol, cationic lipids, PEGylated) and custom liposome formulations.
Liposomes are bilayer (i.e. double-layer), liquid-filled molecules made from phospholipids. Liposome bilayer structure is nearly identical to the cell membranes surrounding the cells in the human body. The phosphate head of phospholipids is hydrophilic (i.e. “water-loving”) and the fatty-acid tails are hydrophobic (i.e. “water-haters”).
When in a water-based solution, the hydrophobic tails distance themselves from the hydrophilic environment. As the tails of the phospholipid turn inward, the hydrophilic heads turn toward outward toward water. This is forms the well-known double-layered membrane. Liposomes are used to protect and deliver siRNA, miRNA and plasmid DNA into specific cells of the body.
Preclinical Research: in vivo toxicology/pharmacology
Toxicology studies are performed after a single, large dose of the test compound is injected and enables optimization of dose route, dose level and maximum tolerated dose. The in vivo transfection reagent or company proprietary test compound is used to determine a safe procedure for administration and can be performed by the appropriate CRO service provider.
The drug development process can be expedited by selecting the appropriate CDX animal model, conducting well designed in vivo studies and following good research practices. Hitting project timelines and staying on budget can be successful by using in vivo CROs. CRO scientists typically have decades worth of preclinical research and project management experience.
Western Blot

Conventional Western Blot technology (immunoblotting) has been utilized for decades, enabling researchers to confirm protein expression levels. Typically, an electrical current moves the proteins through the matrix of an acrylamide gel, thus, separating the proteins based on size. Then, through an archaic membrane transfer method that is highly inefficient, the proteins are transferred to a membrane that is later incubated with an antibody to the target of interest. This is an extremely labor intensive method requiring high sample volumes coupled with innate background issues and low sensitivity.
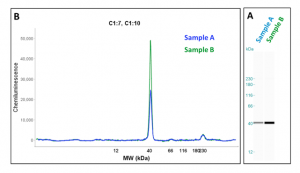
The WES system (by ProteinSimple) is theoretically the same as a conventional western blot, but incorporates modern technology to bypass downfalls of the previous method. This new system enables transfected cellular proteins to be analyzed for changes in expression relative to a non-treated sample. Sensitivity of the WES system is achieved through the use of capillary technology instead of conventional membrane transfer. Once proteins are separated and immobilized within the capillary, reagents such as washes, antibodies and detection enzymes are passed through the capillary. The small capillary size restricts the use of material, thus saving the researcher the cost of reagents.
Stable Cell Line Generation
Exogenous DNA is inserted into plasmid DNA vectors for a variety of reasons, including 1) overexpression for protein production, 2) to study gene specific target gene knockdown, and 3) to create GFP, RFP or luciferase expressing cells to use in cell line derived (CDX) tumor xenograft studies in order to quantitate tumor growth inhibition. An in vivo or in vitro transfection reagent is then used to deliver a payload (e.g. DNA, miRNA, siRNA) to the stable cell line and reduction in fluorescence, bioluminescence or gene expression is quantified.
In Vivo Transfection
Transfection is the process of inserting genetic material, such as siRNA, miRNA, piRNA and plasmid DNA into mammalian cancer cells. The insertion of DNA into a cell enables the expression of the coded protein using the cells typical protein production machinery. The transfected DNA must be transported into the nucleus for effective transfection where the DNA can be transiently expressed or become integrated into the cell’s genomic DNA. With integration, the genetic alteration is passed on to future cells as it divides. Alternatively, RNA insertion into the cytoplasm of a cell is commonly used to down-regulate targeted proteins by stopping translation. There is an array of protocols and techniques available, including lipid transfection, chemical transfection and electroporation.
As a research pipeline progresses from in vitro to in vivo, additional factors must be considered and difficulties will be increased. Special considerations include stability of the DNA or RNA molecule in the presence of serum components that may lead to degradation, specificity of transfection delivery to the desired cells/tissues/organ and efficiency of the delivery agent. The last decade has proven fruitful in the offerings of transfection reagents and transfection kits specifically designed for nucleic acid delivery into a cell line derived xenograft animal model (i.e. RNA, DNA, siRNA, miRNA). There is an array of lipid and non-lipid transfection formulations available, all of which attempt to lower toxicity and increase efficacy.
List of citations and references Altogen Transfection Services and Products
RNAi
RNA interference (RNAi), which is the specific silencing of genes, is a powerful tool in therapeutic and diagnostic development. RNAi is used to identify gene function to determine association with disease phenotypes. Also, gene response pathway changes to chemotherapies or novel compounds is efficiently monitored using high-throughput RNAi screening in mammalian cancer cell lines.
Efficient siRNA design, including chemical modifications that increase stability and target activity, is critical for successful gene knockdown. Initial steps include verifying target gene effects through in vitro transfection experiments since it is a simplified and controlled sample. In vivo transfection reagents are utilized in delivering siRNA, miRNA or DNA to targeted tumor cells to control tumor growth.
Cell Line screens: siRNA and miRNA Library
The ability to perform high-throughput screening of siRNAs or miRNAs is crucial for a complete understanding of response to treatment. All cancers are the result of gaining of a mutation, INDEL or SNP that alters the functionality of the effected gene. Not only are the direct protein products of the affected gene altered, but downstream consequences of the mutation lead to uncontrolled cell proliferation and lack of typical cell death pathway control. With over 30,000 genes to screen in the human genome, CRO services that are capable of transfecting this magnitude of samples are available.
ELISA
An ELISA (enzyme-linked immunosorbent assay) is a plate-based assay able to detect and quantify substances (such as proteins, peptides and antibodies). Briefly, an antigen is immobilized on a solid surface (i.e. bottom of a well) and complexed with an enzyme-linked antibody. The conjugated enzyme activity is assessed by incubation with a substrate that produces a fluorescent product.
This plate based assay enables researchers to track the response of cells transfected with a miRNA or siRNA, or treatment with the company’s test compound. ELISA development offers a quantitative method to determine gene response due to therapeutic targeting, environment influences, toxicity or transfection. The main steps of an ELISA include 1) adsorption to the solid phase plate, 2) addition of test sample and reagents, 3) incubation, 4) washing steps to remove unbound reagents, 5) addition of enzyme labeled reagents, 6) addition of detection system (enzymatic), and 7) quantitative spectrophotometer reading.
Cell Banking
Storage of cancer cell lines in the correct storage buffer and at the correct temperature increases the life span and viability of the cell lines. Due to heavy reliance on control DNA in diagnostic development and target gene expression in therapeutic development, maintaining a reliable cell bank is crucial. Controls (positive and negative) are necessary in transfection studies, and application of controls in storage of cells is of utmost importance. GLP-compliant cell banking systems provide protection and peace of mind for stable cell lines that take months to establish, stem cells, tumor cells and biological materials.
Recombinant DNA Construction
Recombinant DNA technology permits the researcher to sequence genes and then map them to their locations in the genome. Recombinant proteins expressed in cells are used to generate antibody probes for studying protein synthesis. Cloning vectors (represented as yellow in pic below) are DNA molecules able to replicate inside a living cell. A vector is typically a small DNA strand that contains enough genetic information to allow it to be replicated in cells.
Once gene fragments of interest are ligated into the plasmid, bacterial cells are transformed by the recombinant DNA crossing the cell membrane and entering the cytoplasm of the cell. Since the efficiency of transformation of plasmids into bacteria is low, most bacteria cells do not contain the plasmids. However, circular DNA molecules are capable of replication, and plasmids frequently carry genes that benefit their carrier, such as those enabling antibiotic resistance. This provides selection of the positively transfected cells by antibiotic selection; ampicillin resistance gene which can express the enzyme, β-lactamase, to inactivate the antibiotic ampicillin.
Apoptosis Assay
Many diseases, such as cancer, are the result of improperly controlled apoptosis. By using morphological and biochemical modifications to control cell death, cells can self-regulate and genetically ablate themselves under normal operation. Quantification of proteins involved in apoptosis enables researchers to distinguish live cells from early stage apoptotic cells, late stage apoptotic cells and necrotic cells.
Apoptosis is coordinated by members of the caspase family (caspase-3/7) and are activated in response to cell death stimuli. The activated pathway results in dismantling of the cell through proteolysis of cellular proteins. Anti-cancer therapeutics transfected into cells or mice that do not induce apoptosis most likely will lack anti-tumor activity in vivo. For this reason, apoptotic assays are crucial during initial drug screening. Caspase-3/7 levels can be quantified using commercially available fluorescent assays with plate reader results corresponding to the amount of caspase activity in the cells.
Reverse Transfection
Reverse transfection is the process of adding the liposome encapsulated complexes (i.e. transfection reagent plus plasmid DNA/siRNA/miRNA) to a plate well at the same time that cells are being seeded into the plate. This approach is used for large scale RNAi screening studies to enable libraries of pre-plated cargo molecules to be incubated with cultured cells, as well as large scale siRNA library screens using siRNA reverse transfection. Success of these reverse transfections depends on the viability of the cells, the experience of the research technician and the methods employed in the reverse transfection.
microRNA
microRNA is one aspect of the human transcriptome responsible for regulating gene expression. miRNAs control which genes are turned on or off. miRNAs also limit the production of proteins through RNA interference. Research in miRNA involvement in gene expression regulation gained significance when it was found that the human genome contained fewer genes than theorized. Theories suggest miRNAs control 30% of all genes in humans. microRNAs are small units compared to the entirety of the human genome. Endogenous microRNA are a single strand of RNA 20-23 nucleotides in length. Modern technologies enable miRNA to be isolated, amplified and sequenced for quantitation.
Visit Altogen’s Transfection Resource: methods, protocols, stable and transient transfection, RNAi and more.